AI-driven medical imaging has emerged as a transformative force in modern healthcare, empowering clinicians to deliver more accurate, efficient, and personalized diagnostic and therapeutic strategies. By automatically analyzing CT, MRI, ultrasound, and other imaging modalities, AI systems can identify subtle patterns, precisely segment anatomical structures, and support clinical decision-making with enhanced accuracy and consistency. These advancements not only improve diagnostic performance but also streamline clinical workflows and ultimately elevate the overall quality of patient care.
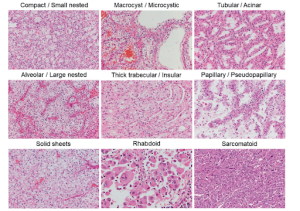
Among the many tasks in medical image analysis, kidney and kidney tumor segmentation are pivotal for the management of renal diseases, particularly renal cell carcinoma. Precise delineation of kidneys and tumors is essential for quantitative tumor assessment, treatment planning, surgical navigation, postoperative monitoring, and radiomics research. Accurate segmentation enables clinicians to reliably estimate tumor burden, evaluate tumor–organ spatial relationships, and facilitate nephron-sparing surgical strategies, all of which directly influence patient outcomes. Given that manual segmentation is labor-intensive and prone to inter- and intra-observer variability, the development of automated, robust, and reliable segmentation methods has become increasingly critical for both routine clinical practice and large-scale research.
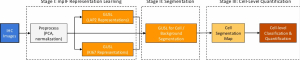
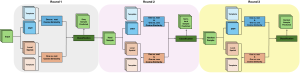
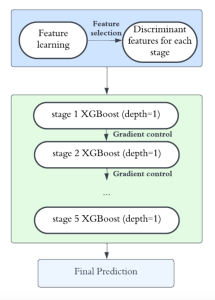
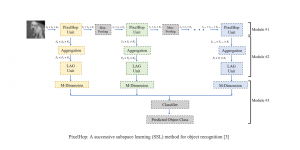
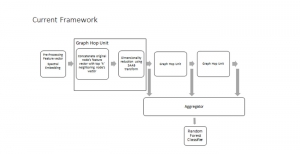
To address these limitations, we propose 3D-Cube Multi-Stage Green U-shaped Learning (GUSL), a novel multi-stage feed-forward machine learning framework for 3D medical image segmentation without backpropagation. GUSL is designed to be computationally efficient, interpretable, and environmentally sustainable, while maintaining competitive segmentation performance.
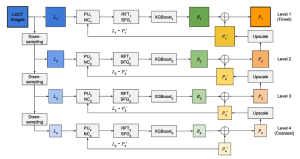
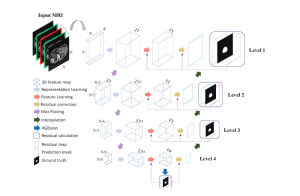
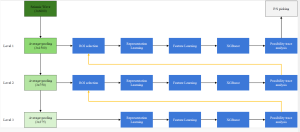
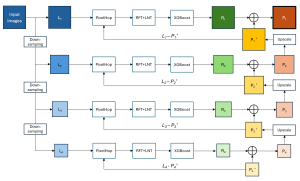
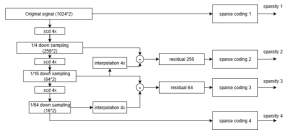
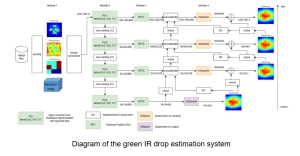
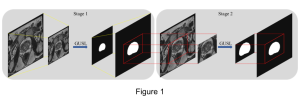
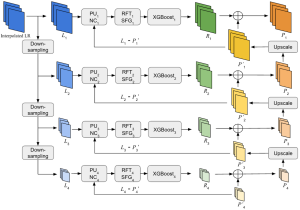
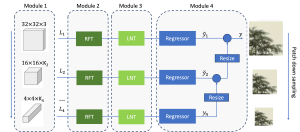
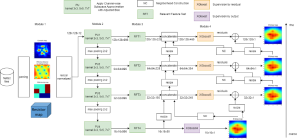
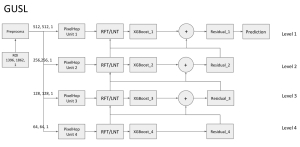
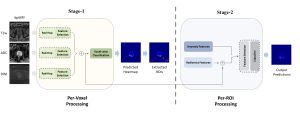
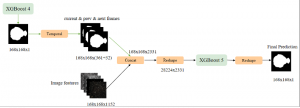
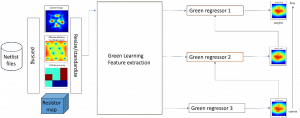
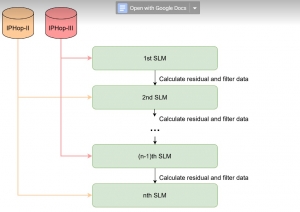
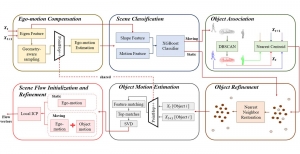
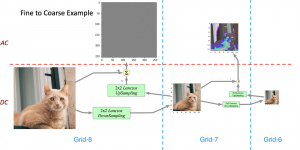
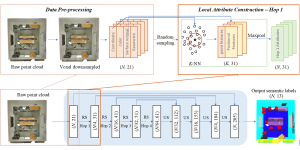
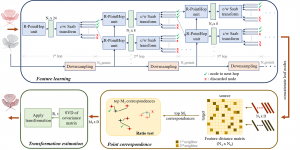
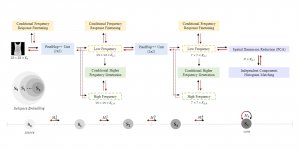
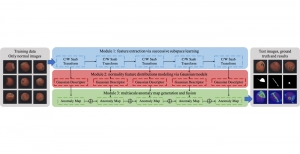
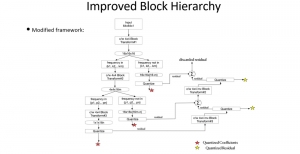
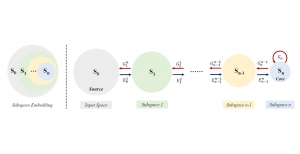
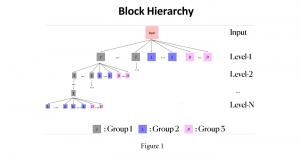
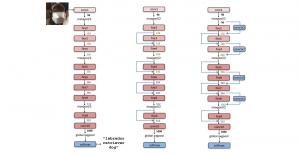
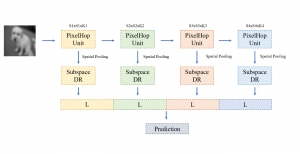
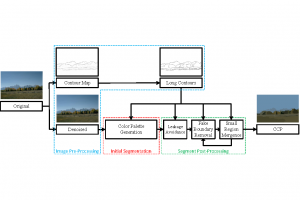
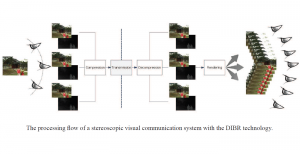
The proposed framework adopts a cascaded multi-stage segmentation strategy tailored to different anatomical tasks. As illustrated in Figure 1, distinct stages are designed for coarse-to-fine segmentation. First, the original CT volume is downsampled to a lower resolution, enabling efficient coarse localization of the kidney with reduced computational complexity. This low-resolution stage provides an approximate spatial position of the kidney, as highlighted in the red box. The kidney Region-of-Interest (ROI) is then cropped from the original-resolution image to preserve structural details for subsequent refinement.
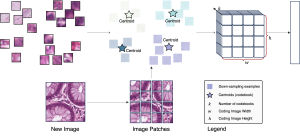
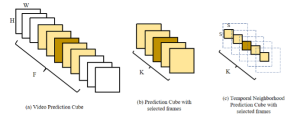
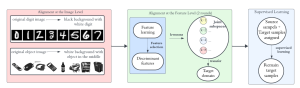
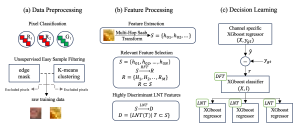
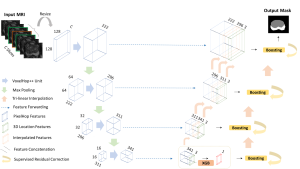
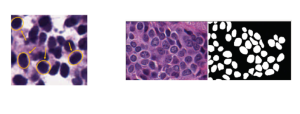
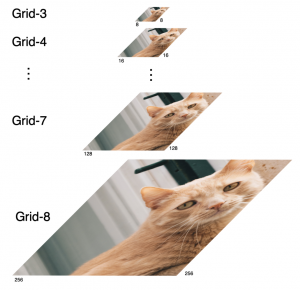
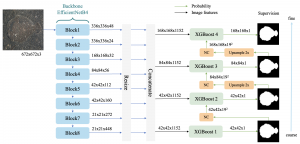
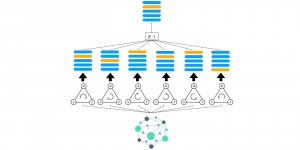
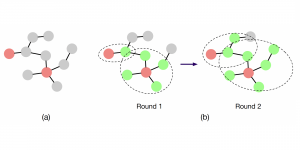
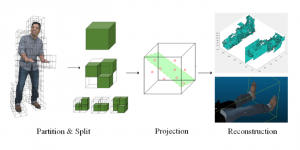
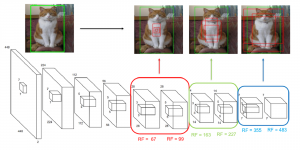
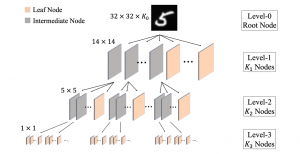
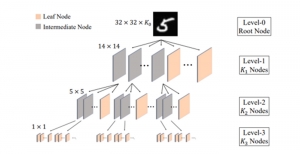
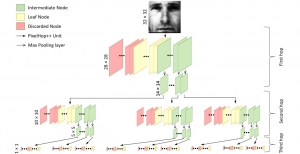
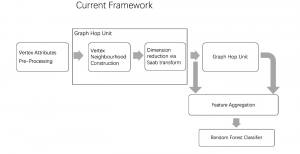
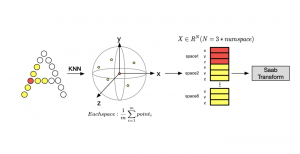
Within the cropped kidney ROI, detailed segmentation of the kidney, mass, and tumor structures is performed. Due to severe class imbalance in mass and tumor segmentation, we further partition each kidney ROI into 64×64×64 3D cubes, as shown in Figure 2. This cube-based strategy effectively reduces redundant negative samples and facilitates targeted data augmentation for minority classes, improving learning efficiency and robustness.
Extensive experiments on the KiTS23 dataset demonstrate that GUSL achieves state-of-the-art performance in kidney, mass, and tumor segmentation. Beyond segmentation accuracy, GUSL consistently yields substantial reductions in model size, computational complexity (FLOPs), energy consumption, and carbon footprint, underscoring its clear advantages over conventional deep learning approaches.